Domains
The following domain information was found using PFAM and SMART (1, 2).
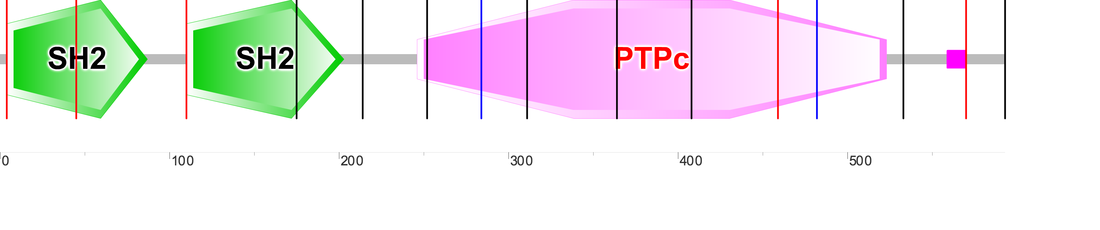
PTPN11 contains two distinct protein domains:
1) Tyrosine (amino acid Y) protein phosphatase is part of a group of enzymes which remove phosphates from phosphorylated tyrosine residues on other proteins. Tyrosine phosphorylation is a common type of post-translation protein modification that can give proteins more specificity, potential protein interactions, and also alter enzymatic reactions within the cell. Y_phosphatase acts as a regulator of these functions by maintaining a stable level of phosphorylated tyrosine residues within a cell.
2) The SH2 (Src Homology 2) domain is a highly conserved protein domain among higher organisms known for its indispensability in intracellular signal transduction pathways. SH2 domains allow proteins containing those domains to dock to phosphorylated tyrosine residues on other proteins. For this reason, SH2 domains are generally found within adapter proteins, assisting in the signal transduction of receptor tyrosine kinase pathways (3).
The following domain information was found using PFAM and SMART (1, 2).
PTPN11 contains two distinct protein domains:
1) Tyrosine (amino acid Y) protein phosphatase is part of a group of enzymes which remove phosphates from phosphorylated tyrosine residues on other proteins. Tyrosine phosphorylation is a common type of post-translation protein modification that can give proteins more specificity, potential protein interactions, and also alter enzymatic reactions within the cell. Y_phosphatase acts as a regulator of these functions by maintaining a stable level of phosphorylated tyrosine residues within a cell.
2) The SH2 (Src Homology 2) domain is a highly conserved protein domain among higher organisms known for its indispensability in intracellular signal transduction pathways. SH2 domains allow proteins containing those domains to dock to phosphorylated tyrosine residues on other proteins. For this reason, SH2 domains are generally found within adapter proteins, assisting in the signal transduction of receptor tyrosine kinase pathways (3).
Figure 1: Domains generated by SMART (3)
Figure 2: Domains generated by PFAM (2)
Motifs
DNA sequence motifs are recurring patterns that are predicted to have a biological function within an organisms genome due to their presence in other organisms. They often suggest sequence-specific binding sites for regulatory proteins like transcription factors. Furthermore, some motifs may take effect in processes at the RNA level that regulate amino acid synthesis (4).
Interpreting DNA Motifs:
Using MEME, motifs 1-3 were generated based on PTPN11 DNA sequence data. The numbers on the x-axis of the motif represent the base pair of that specific motif while the y-axis indicates the likelihood of the represented base. For example, at base pair 13 there are two potential bases: G (guanine) and A (adenine). Since the letter G is larger than the letter A, the liklihood of the base being a guanine is greater (5).
DNA sequence motifs are recurring patterns that are predicted to have a biological function within an organisms genome due to their presence in other organisms. They often suggest sequence-specific binding sites for regulatory proteins like transcription factors. Furthermore, some motifs may take effect in processes at the RNA level that regulate amino acid synthesis (4).
Interpreting DNA Motifs:
Using MEME, motifs 1-3 were generated based on PTPN11 DNA sequence data. The numbers on the x-axis of the motif represent the base pair of that specific motif while the y-axis indicates the likelihood of the represented base. For example, at base pair 13 there are two potential bases: G (guanine) and A (adenine). Since the letter G is larger than the letter A, the liklihood of the base being a guanine is greater (5).
Figure 1: Motif 1

Figure 2: Motif 2
Figure 3: Motif 3
|
Chemical Genomics
Closely related to pharmaceuticals, chemical genomics is the study of how small molecules interact with proteins or other biochemical pathways such as signal transduction. These small molecules are important because their chemical interactions with biochemical molecules can be used to cure disease by inhibiting, altering, or promoting certain proteins. Generally used for the discovery of novel drugs to treat diseases, almost every person on earth has benefited from these small molecules. For example, by taking antihistamines for allergies, you are inserting chemicals into your body that inhibit histamines that promote an allergic response (6). Online databases such as Pubchem, Chembank, and emolecules are used by researchers to find and analyze chemicals that alter specific proteins or genes (7, 8, 9). |
Figure 4: Antihistamine example (10)
|
Have molecules been found that affect SHP2?
According to the Pubchem online database, there are numerous experimental SHP2 inhibitors that have been found to counteract the overactivity of mutated SHP2 (11). The drugs currently being tested function by binding to the catalytic site of SHP2, inhibiting its phosphatase ability and reducing incorrect signal transduction which causes the symptoms of NS. However, the catalytic site of SHP2 closely matches the catalytic site of SHP1 which functions normally in patients with NS. Another area of concern is the prevalence of SH2 motifs in other proteins that could also be affected by these drugs. The objective of current research is to develop a drug with greater specificity to the SHP2 catalytic site in order to remove off-target effects (12).
According to the Pubchem online database, there are numerous experimental SHP2 inhibitors that have been found to counteract the overactivity of mutated SHP2 (11). The drugs currently being tested function by binding to the catalytic site of SHP2, inhibiting its phosphatase ability and reducing incorrect signal transduction which causes the symptoms of NS. However, the catalytic site of SHP2 closely matches the catalytic site of SHP1 which functions normally in patients with NS. Another area of concern is the prevalence of SH2 motifs in other proteins that could also be affected by these drugs. The objective of current research is to develop a drug with greater specificity to the SHP2 catalytic site in order to remove off-target effects (12).
Protein Interactions
What is a protein interaction?
In biological systems, proteins are the driving force behind cell homeostasis, metabolism, and signal transduction. A protein interaction is simply an indication that one protein physically binds to and/or alters another protein. These interactions can promote or inhibit a protein's ability to carry out its function and/or allow a protein to be used for another function.
What does PTPN11 interact with?
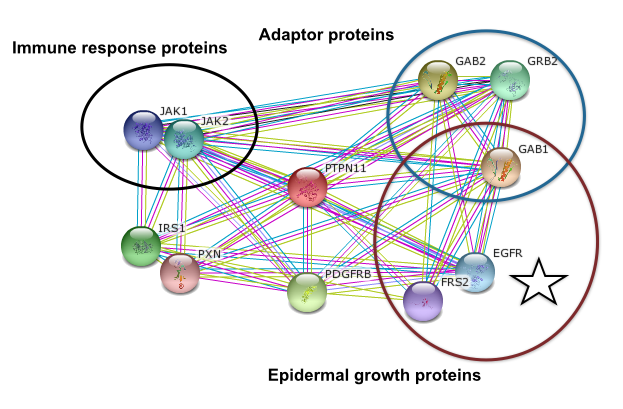
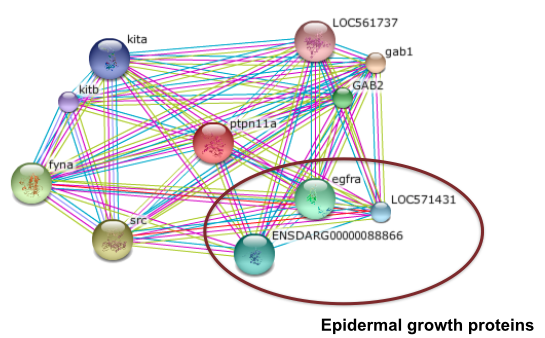
As a signal transduction molecule, PTPN11 has many protein interactions that are strongly suggested by experimental or compuatational evidence. In humans, PTPN11 interacts with proteins involved with epidermal growth, immune response, insulin response, and numerous adaptor proteins that connect it to other signal transduction pathways. In zebrafish, PTPN11 interacts with adaptor proteins but, more importantly, it interacts with epidermal growth proteins (13).
What does PTPN11 interact with?
As a signal transduction molecule, PTPN11 has many protein interactions that are strongly suggested by experimental or compuatational evidence. In humans, PTPN11 interacts with proteins involved with epidermal growth, immune response, insulin response, and numerous adaptor proteins that connect it to other signal transduction pathways. In zebrafish, PTPN11 interacts with adaptor proteins but, more importantly, it interacts with epidermal growth proteins (13).
Figure 5: Human PTPN11 interaction network (13)
Figure 6: Zebrafish PTPN11 interaction network (13)
Why are these protein interactions important?
These protein interactions provide a basis for understanding the cellular effect of a mutated protein. Since PTPN11 interacts with epidermal growth factor receptor (EGFR) in humans, it is suggested that this interaction may cause some of the symptoms of people with NS since they usually have unusual skin growth. Furthermore, since this interaction is also expressed in zebrafish, zebrafish can be used for tests involving EGFR. Lastly, knowing which proteins interact with PTPN11 allows us to make informed targets for chemical compounds (drugs) to alleviate the symptoms of NS.
These protein interactions provide a basis for understanding the cellular effect of a mutated protein. Since PTPN11 interacts with epidermal growth factor receptor (EGFR) in humans, it is suggested that this interaction may cause some of the symptoms of people with NS since they usually have unusual skin growth. Furthermore, since this interaction is also expressed in zebrafish, zebrafish can be used for tests involving EGFR. Lastly, knowing which proteins interact with PTPN11 allows us to make informed targets for chemical compounds (drugs) to alleviate the symptoms of NS.
Protein Modifications
|
What is a protein modification?
|
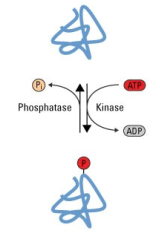
Figure 7: Protein phosphorylation (14)
|
|
Protein modifications are alterations made to proteins during or after their
translation. These alterations usually involve chemical modifications to individual amino acids that cannot be acheived by translation and greatly increase the amount of functions a single type of protein can have. While there are many different types of protein modifications, phosphorylation is the most common and is a large focus of research in the proteomic field. |
What kind of translation modifications does PTPN11 have and why are they important?
Due to its involvement in many diseases, PTPN11 is a widely studied protein; thus, post-translational modifications of PTPN11 were easily obtained from the Uniprot online database.
PTPN11 has the following residues post-translationally phosphorylated: Tyr-62, Tyr-66, Tyr-546, Tyr-546, and Tyr-584. Phosphorylation of Tyrosine residues 546 and 584 allows PTPN11 to bind to GRB2 adaptor protein, promoting signal propagation. These residues may prove to be good drug targets in order to inhibit overactive PTPN11 in patients with NS.
Furthermore, it was found that residue 2 has an addition of acetyl-threonine. However, since this modification is seen on many proteins due to its role as a cell localization factor, it is not recommended to be a drug target (15).
PTPN11 has the following residues post-translationally phosphorylated: Tyr-62, Tyr-66, Tyr-546, Tyr-546, and Tyr-584. Phosphorylation of Tyrosine residues 546 and 584 allows PTPN11 to bind to GRB2 adaptor protein, promoting signal propagation. These residues may prove to be good drug targets in order to inhibit overactive PTPN11 in patients with NS.
Furthermore, it was found that residue 2 has an addition of acetyl-threonine. However, since this modification is seen on many proteins due to its role as a cell localization factor, it is not recommended to be a drug target (15).
References
1) "Pfam: Home Page." Pfam: Home Page. EMBL/EBI, n.d. Web. 27 Mar. 2015.
2) "Select Your Default SMART Mode." SMART: Main Page. N.p., n.d. Web. 27 Mar. 2015.
3) "Domain Architecture Analysis." SMART: Sequence Analysis Results for ENSP00000340944. N.p., n.d. Web. 27 Mar. 2015.
4) D'haeseleer, Patric. "What Are DNA Sequence Motifs?" Nature.com. Nature Publishing Group, n.d. Web. 27 Mar. 2015.
5) "The MEME Suite." MEME. NBCR, n.d. Web. 27 Mar. 2015.
6) Roemer, Terry, Julian Davies, Guri Giaever, and Corey Nislow. "Bugs, Drugs and Chemical Genomics." Nature.com. NPG, 15 Dec. 2011. Web. 27 Mar. 2015.
7) "The PubChem Project." Whats New in PubChem RSS. NIH, n.d. Web. 27 Mar. 2015.
8) "ChemBank." Home. ChemAxon, n.d. Web. 27 Mar. 2015.
9) "Chemical Structure Drawing Search - EMolecules." EMolecules. N.p., n.d. Web. 27 Mar. 2015.
10) "Benadryl Allergy Antihistamine Tablets." CVS.com. N.p., n.d. Web. 27 Mar. 2015.
11) "PubChem Search." PubChem. N.p., n.d. Web. 27 Mar. 2015.
12) Hartman, Zachary. "Characterizing SHP2 as a Novel Therapeutic Target in Breast Cancer." Characterizing SHP2 as a Novel Therapeutic Target in Breast Cancer. N.p., Feb. 2013. Web. 27 Mar. 2015.
13) "STRING: Functional Protein Association Networks." STRING: Functional Protein Association Networks. SIB, n.d. Web. 06 May 2015.
14) "Phosphorylation." Phosphorylation. N.p., n.d. Web. 16 May 2015.
15) "Tyrosine-protein Phosphatase Non-receptor Type 11." Uniprot. EMBL-EBI, n.d. Web. 16 May 2015.
1) "Pfam: Home Page." Pfam: Home Page. EMBL/EBI, n.d. Web. 27 Mar. 2015.
2) "Select Your Default SMART Mode." SMART: Main Page. N.p., n.d. Web. 27 Mar. 2015.
3) "Domain Architecture Analysis." SMART: Sequence Analysis Results for ENSP00000340944. N.p., n.d. Web. 27 Mar. 2015.
4) D'haeseleer, Patric. "What Are DNA Sequence Motifs?" Nature.com. Nature Publishing Group, n.d. Web. 27 Mar. 2015.
5) "The MEME Suite." MEME. NBCR, n.d. Web. 27 Mar. 2015.
6) Roemer, Terry, Julian Davies, Guri Giaever, and Corey Nislow. "Bugs, Drugs and Chemical Genomics." Nature.com. NPG, 15 Dec. 2011. Web. 27 Mar. 2015.
7) "The PubChem Project." Whats New in PubChem RSS. NIH, n.d. Web. 27 Mar. 2015.
8) "ChemBank." Home. ChemAxon, n.d. Web. 27 Mar. 2015.
9) "Chemical Structure Drawing Search - EMolecules." EMolecules. N.p., n.d. Web. 27 Mar. 2015.
10) "Benadryl Allergy Antihistamine Tablets." CVS.com. N.p., n.d. Web. 27 Mar. 2015.
11) "PubChem Search." PubChem. N.p., n.d. Web. 27 Mar. 2015.
12) Hartman, Zachary. "Characterizing SHP2 as a Novel Therapeutic Target in Breast Cancer." Characterizing SHP2 as a Novel Therapeutic Target in Breast Cancer. N.p., Feb. 2013. Web. 27 Mar. 2015.
13) "STRING: Functional Protein Association Networks." STRING: Functional Protein Association Networks. SIB, n.d. Web. 06 May 2015.
14) "Phosphorylation." Phosphorylation. N.p., n.d. Web. 16 May 2015.
15) "Tyrosine-protein Phosphatase Non-receptor Type 11." Uniprot. EMBL-EBI, n.d. Web. 16 May 2015.