Introduction
Also known as Leopard Syndrome, Noonan syndrome (NS) is a developmental disease that is characterized by short stature, unusual facial characteristics, heart defects, skeletal malformations and numerous other physical defects. However, these physical characteristics can vary significantly between individuals due to the various molecular causes of NS (1). PTPN11/SHP2 is the most commonly mutated protein in people with NS and while there are many treatments that target various symptoms caused by NS, there are currently limited options available that target PTPN11/SHP2.
According to the string database, PTPN11 interacts with numerous proteins involved with signal transduction including epidermal growth factor receptor (EGFR) (2). EGFR is a protein that promotes skin growth upon stimulation of certain growth factors. Since many people diagnosed with Noonan syndrome experience higher rates of cancer and have excessive amounts of skin in some locations, it is hypothesized that PTPN11 regulates proteins involved with skin development (1). The primary goal of the following aims is to discover proteins involved in embryonic skin development and learn how to effectively manipulate them on a chemogenomic level.
According to the string database, PTPN11 interacts with numerous proteins involved with signal transduction including epidermal growth factor receptor (EGFR) (2). EGFR is a protein that promotes skin growth upon stimulation of certain growth factors. Since many people diagnosed with Noonan syndrome experience higher rates of cancer and have excessive amounts of skin in some locations, it is hypothesized that PTPN11 regulates proteins involved with skin development (1). The primary goal of the following aims is to discover proteins involved in embryonic skin development and learn how to effectively manipulate them on a chemogenomic level.
Specific Aims
Aim 1 - Determine wildtype and mutant protein interactions controlling skin development
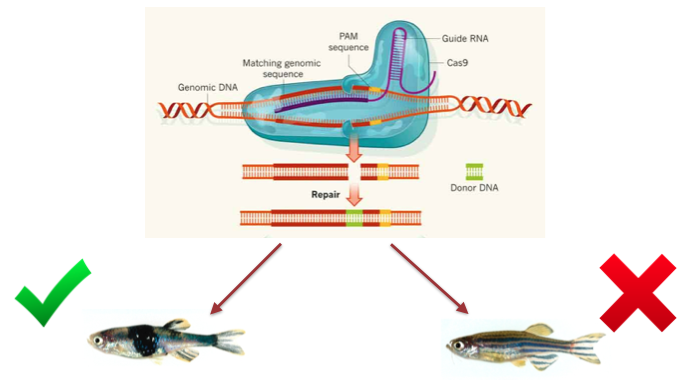
The first aim is focused on discovering proteins that interact with PTPN11 and EGFR and are involved with skin development. In order to do so, transgenic zebrafish with mutated PTPN11 will be generated via CRISPR. Zebrafish will be used because of their fast generation time, large brood size, and transparency during development. Once the transgenic strains are generated, tandem affinity purification (TAP) will be used to elucidate proteins that both EGFR and PTPN11 interact with using both the proteins as bait in seperate assays. However, association with these proteins does not provide confirmation that the protein actually controls skin development. However, In order to confirm that the interacting proteins do have a role in skin development, they will be knocked-out in zebrafish via CRISPR and the phenotypic effects will be monitored.
Aim 2 - Identify compounds that inhibit WT and/or mutant PTPN11 interacting proteins
The goal of the second aim is to find chemical compounds (drugs) that have an inhibitory effect on the interactions of potential interacting proteins and PTPN11 or EGFR. Initially, chemically active compounds will be found based on the contents of Pubchem database (5). If there are insufficient amounts of active compounds available, we will conduct a chemogenomic assay to screen for potential inhibitors of wildtype and mutant proteins interactions.
Figure 2: Chemogenomic assay (6)
Aim 3: Test active compound efficacy using phophoproteomics
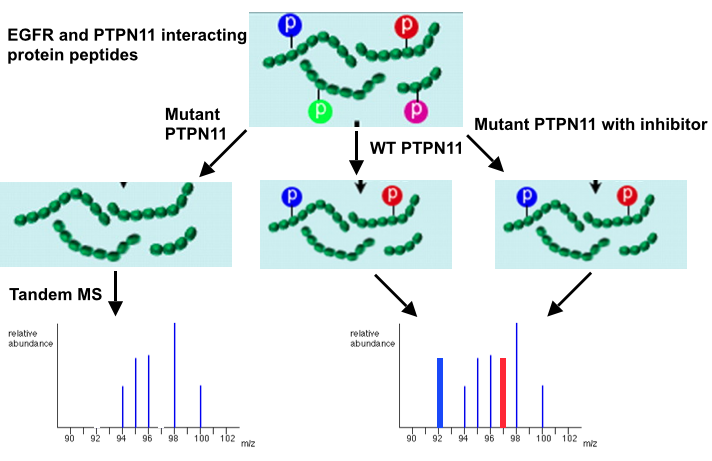
An in vitro phosphoproteomic assay using tandem mass spectrometry will be used in order to test the efficacy of discovered active compounds. First, B normally phosphorylated protein will be exposed to wildtype and mutant PTPN11 in the presence of active compounds. After exposure, these proteins will be subjected to tandem mass spectrometry to determine the how much of the protein is still phosphorylated. Since PTPN11 removes phosphates from proteins, and mutated PTPN11 is overactive, it is expected that in trials with mutant PTPN11, all phosphate will be eliminated from the protein. However, under the effects of certain active compounds, it is expected that the amount of phosphorylated residues on the protein after mutant PTPN11 exposure will resemble that of a protein exposed to wildtype PTPN11, with some phosphorylated residues remaining.
An in vitro phosphoproteomic assay using tandem mass spectrometry will be used in order to test the efficacy of discovered active compounds. First, B normally phosphorylated protein will be exposed to wildtype and mutant PTPN11 in the presence of active compounds. After exposure, these proteins will be subjected to tandem mass spectrometry to determine the how much of the protein is still phosphorylated. Since PTPN11 removes phosphates from proteins, and mutated PTPN11 is overactive, it is expected that in trials with mutant PTPN11, all phosphate will be eliminated from the protein. However, under the effects of certain active compounds, it is expected that the amount of phosphorylated residues on the protein after mutant PTPN11 exposure will resemble that of a protein exposed to wildtype PTPN11, with some phosphorylated residues remaining.
Figure 3: Phosphoproteomic experimental design (7)
Future Directions
If active compounds are found using this assay, we will first move on to in vivo testing in zebrafish in order to evaluate side effects incurred from potentially active compounds. It is expected that there may have to be many adjustments to be made on active compounds in order to calibrate their strength and reduce their off-target effects in live tissue. If active compounds are found to be effective in zebrafish with minimal side effects, testing will continue on an animal more similar to humans, most likely mice.
In regards to NS research as a whole, a technique that targets proteins downstream of PTPN11 with multiple sets of compounds may be more effective than targeting only PTPN11 at the source. Since some PTPN11-based pathways may remain unaffected by mutated PTPN11, targeting specific downstream pathways reduces the chance of interfering with pathways that are still functional. Furthermore, the knowledge discovered by these aims may prove beneficial not only to people with NS, but others that suffer a variety of cancers since PTPN11 plays such a pivotal role in these types of diseases as well (1).
In regards to NS research as a whole, a technique that targets proteins downstream of PTPN11 with multiple sets of compounds may be more effective than targeting only PTPN11 at the source. Since some PTPN11-based pathways may remain unaffected by mutated PTPN11, targeting specific downstream pathways reduces the chance of interfering with pathways that are still functional. Furthermore, the knowledge discovered by these aims may prove beneficial not only to people with NS, but others that suffer a variety of cancers since PTPN11 plays such a pivotal role in these types of diseases as well (1).
| ptpn11_final_talk.pdf | |
| File Size: | 2401 kb |
| File Type: | |
References
1) "Noonan Syndrome." Genetics Home Reference. NIH, Mar. 2011. Web. 27 Mar. 2015.
2) "STRING: Functional Protein Association Networks." STRING: Functional Protein Association Networks. SIB, n.d. Web. 06 May 2015.
3) "CRISPR/Cas9 Genome Editing." CRISPR Cas9 Genome Editing. N.p., n.d. Web. 16 May 2015.
4) "Zebra Fish Ipilimumab Las Vegas Dermatology." Helios Dermatology . N.p., n.d. Web. 16 May 2015.
5) "The PubChem Project." Whats New in PubChem RSS. NIH, n.d. Web. 27 Mar. 2015.
6) "96-Well Treated Microplates: U- Bottom [50063] | EdgeBio." Edgebio. N.p., n.d. Web. 16 May 2015.
7) "The Mass Spectrometer - How It Works." Chemguide. N.p., n.d. Web. 16 May 2015.
2) "STRING: Functional Protein Association Networks." STRING: Functional Protein Association Networks. SIB, n.d. Web. 06 May 2015.
3) "CRISPR/Cas9 Genome Editing." CRISPR Cas9 Genome Editing. N.p., n.d. Web. 16 May 2015.
4) "Zebra Fish Ipilimumab Las Vegas Dermatology." Helios Dermatology . N.p., n.d. Web. 16 May 2015.
5) "The PubChem Project." Whats New in PubChem RSS. NIH, n.d. Web. 27 Mar. 2015.
6) "96-Well Treated Microplates: U- Bottom [50063] | EdgeBio." Edgebio. N.p., n.d. Web. 16 May 2015.
7) "The Mass Spectrometer - How It Works." Chemguide. N.p., n.d. Web. 16 May 2015.