What is PTPN11?
Protein tyrosine phosphatase non-receptor type 11 (PTPN11) is the most commonly mutated gene for patients with Noonan Syndrome (NS). PTPN11 encodes for the protein SHP-2 which controls the activation of the RAS/MAPK pathway which plays a large role in development of the heart, bones, blood cells, and other tissues. In humans, the cytogenetic location of PTPN11 is 12q24. As a member of the PTP gene family, PTPN11 assists the cell in removing phosphate groups from other proteins, altering their function. Through this phosphatase function, proteins created from the PTP gene family are very important to regulating inter-cellular communication through signal transduction and has a large role in controlling cell growth in response to hormones (1).
Human Bioinformatic information
Accession Number - NP_002825.3
FASTA (2)
Protein tyrosine phosphatase non-receptor type 11 (PTPN11) is the most commonly mutated gene for patients with Noonan Syndrome (NS). PTPN11 encodes for the protein SHP-2 which controls the activation of the RAS/MAPK pathway which plays a large role in development of the heart, bones, blood cells, and other tissues. In humans, the cytogenetic location of PTPN11 is 12q24. As a member of the PTP gene family, PTPN11 assists the cell in removing phosphate groups from other proteins, altering their function. Through this phosphatase function, proteins created from the PTP gene family are very important to regulating inter-cellular communication through signal transduction and has a large role in controlling cell growth in response to hormones (1).
Human Bioinformatic information
Accession Number - NP_002825.3
FASTA (2)
Gene Ontology
Known as the Gene Ontology Consortium, researchers have classified into three categories the thousands of gene products synthesized by model organisms. These three categories are cellular component, biological process, and molecular function. The cellular component category describes where in a cell that a specific gene localizes. Biological processes describes the series of events caused by the presence of a gene. Lastly, molecular function describes the activities of the gene at a molecular level such as protein interactions or other chemical activities (3).
Known as the Gene Ontology Consortium, researchers have classified into three categories the thousands of gene products synthesized by model organisms. These three categories are cellular component, biological process, and molecular function. The cellular component category describes where in a cell that a specific gene localizes. Biological processes describes the series of events caused by the presence of a gene. Lastly, molecular function describes the activities of the gene at a molecular level such as protein interactions or other chemical activities (3).
|
Molecular function
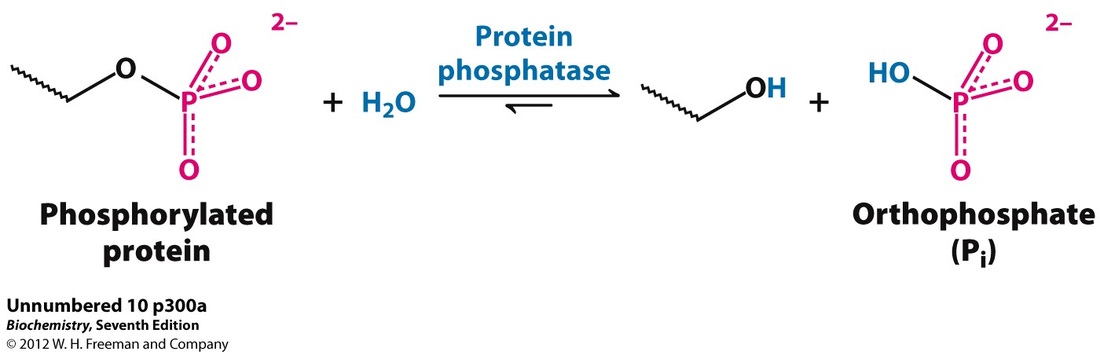
Protein phosphatase (GO:0004726) (3) *There are other molecular functions, however, they are unrelated to the pathogenesis of NS |
Figure 1: Chemical process of a phosphatase (4)
|
|
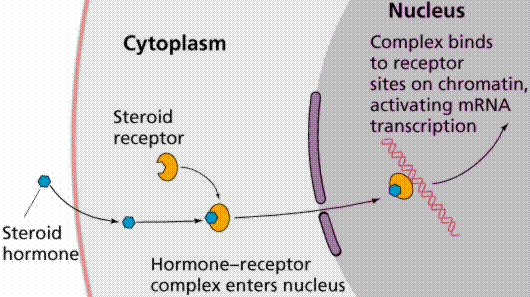
Figure 2: Hormonal pathway (5)
|
Biological process:
Downregulates secretion of: Growth hormones (GO:0060125) Cortisol (GO:0051463) Steroids (GO:2000832) Glucocorticoids (GO:2000850) (3) |
|
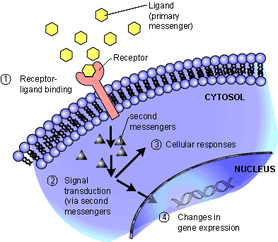
Cellular component:
Growth hormone receptor complex (GO:0070195) Intracellular cyclic nucleotide activated cation channel complex (GO:0017071) Ectoplasm (GO:0043265) Subapical complex (GO:0035003) (3) |
Figure 3: Signal Transduction (6)
|
Analysis:
In regards to Noonan Syndrome, PTPN11 plays a major role in the development of organisms in response to endocrinol signaling. This is shown by the its biological process ontology which shows its role in the regulation of hormones and steroids. Furthermore, in the cellular component section, it is shown to localize in areas of the cell near the cell membrane, where important steps of endocrinol signaling take place. Lastly, molecular function describes its role in these processes at the given location as a protein phosphatase.Given these associations, it is logical that malfunctions in PTPN11 lead to improper growth of skin and overall stature.
In regards to Noonan Syndrome, PTPN11 plays a major role in the development of organisms in response to endocrinol signaling. This is shown by the its biological process ontology which shows its role in the regulation of hormones and steroids. Furthermore, in the cellular component section, it is shown to localize in areas of the cell near the cell membrane, where important steps of endocrinol signaling take place. Lastly, molecular function describes its role in these processes at the given location as a protein phosphatase.Given these associations, it is logical that malfunctions in PTPN11 lead to improper growth of skin and overall stature.
|
Model Organisms: Mutant Phenotypes and Mutagenesis
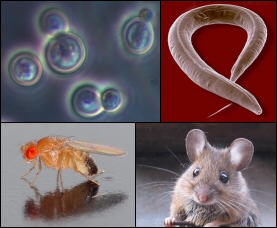
What is a model organism? Model organisms are species that have been found to be extremely useful in biological scientific experiments due to certain advantages that aid in research. These organisms tend to have quick generation periods, are cheap and easy to maintain, and have physiological similarities with humans. Furthermore, due to the extensive use of these organisms in the past, reseachers have extensive background knowledge of their model species (8). Common model organisms include mice, zebrafish, fruit flies, nematodes, arabadopsis, yeast, and E. coli. |
Figure 4: Model Organisms (7)
|
What model organisms can be used for NS research?
In the scientific field, researchers studying NS have used three transgenic model organisms: Zebrafish, Drosophila, and Mice.
1) Zebrafish with effective NS were found to display gain-of-function phenotypes that are very similar to human phenotypes. These include reduced stature, absence of swim bladder, and the abnormal development of eyes, heart, and head. Using zebrafish is advantageous due to their transparent nature during development, the rate at which they grow, and their fairly large brood size (9).
2) Drosophila with effective NS also display gain-of-function phenotypes. However, it is not clear how these relate to the human phenotypes. Furthermore, due to their equivalent PTPN11 gene only being 66% identical, it is likely that the protein serves different functions in drosophila than in humans. Advantages drosophila have over these other model organisms include their rapid generation time and massive brood size (10).
3) Mice with effective NS display the most homologous phenotypic effects of NS compared to humans. Easily observed phenotypes include cardiac defects, reduced stature, shortened lifespan, and unusual facial characteristics. While the phenotypes of mice closely match humans and are the best comparison of the three, their slow generation time and relatively small progeny size make them disadvantageous for many experiments (11).
How do researchers create these transgenic model organisms?
Scientists use the following techniques to create organisms that can be used to model their disease of choice:
In the scientific field, researchers studying NS have used three transgenic model organisms: Zebrafish, Drosophila, and Mice.
1) Zebrafish with effective NS were found to display gain-of-function phenotypes that are very similar to human phenotypes. These include reduced stature, absence of swim bladder, and the abnormal development of eyes, heart, and head. Using zebrafish is advantageous due to their transparent nature during development, the rate at which they grow, and their fairly large brood size (9).
2) Drosophila with effective NS also display gain-of-function phenotypes. However, it is not clear how these relate to the human phenotypes. Furthermore, due to their equivalent PTPN11 gene only being 66% identical, it is likely that the protein serves different functions in drosophila than in humans. Advantages drosophila have over these other model organisms include their rapid generation time and massive brood size (10).
3) Mice with effective NS display the most homologous phenotypic effects of NS compared to humans. Easily observed phenotypes include cardiac defects, reduced stature, shortened lifespan, and unusual facial characteristics. While the phenotypes of mice closely match humans and are the best comparison of the three, their slow generation time and relatively small progeny size make them disadvantageous for many experiments (11).
How do researchers create these transgenic model organisms?
Scientists use the following techniques to create organisms that can be used to model their disease of choice:
|
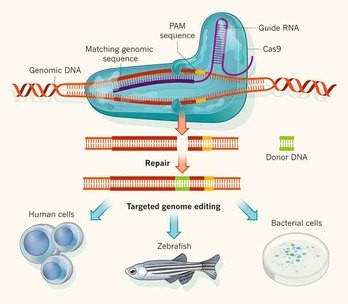
Figure 5: CRISPR/Cas9 mechanism (12)
|
CRISPR/Cas9 system: The CRISPR/Cas system is a bacterial immune system that confers resistance to viruses and foreign DNA, giving a cell a form of acquired immunity. CRISPR consists of spacers that can be transcribed into RNA that recognize foreign genetic material inside a cell. Cas9 is a nuclease (type of enzyme) that recognizes CRISPR RNA conjugated with foreign genetic material and degrades them. Recently, the CRISPR/Cas9 system has been used for gene manipulation in many model organisms for its selective gene modifying abilities. Furthermore, due to CRISPR's dependence on Cas9 for DNA degradation, this gene manipulation can be induced at a time preferable to researchers (13). |
|
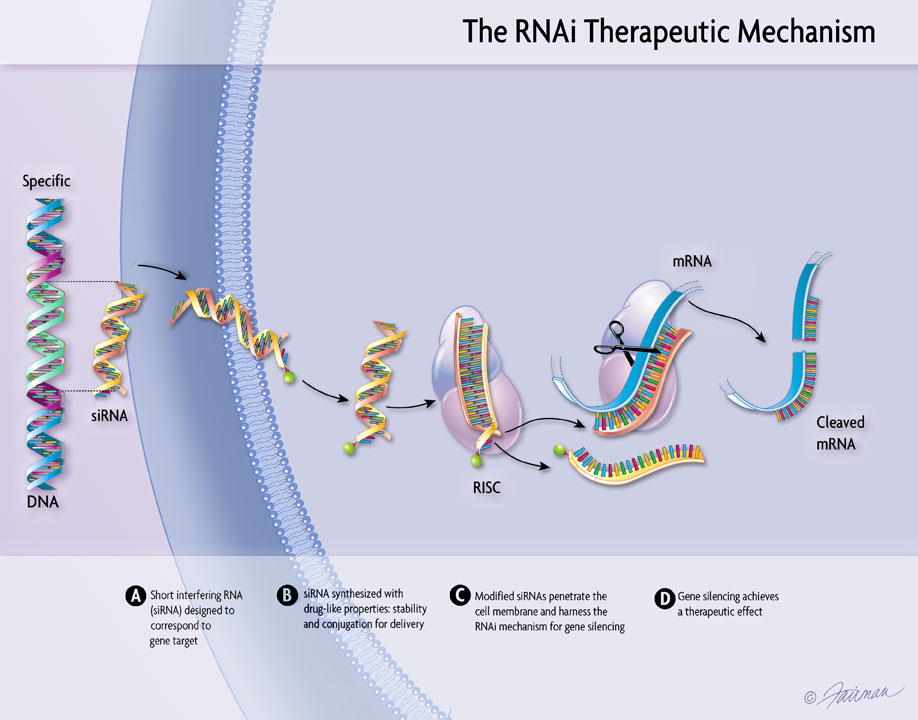
RNAi: This technique works by introducing small interfering RNA (siRNA) or micro RNA (miRNA) sequences that bind to select mRNA transcripts being expressed by a cell. By binding to the expressed mRNA, siRNA/miRNA inhibit production of the specific protein the mRNA codes for. Furthermore, conjugation of siRNA/mRNA also induces its own degradation by an RNA-induced silencing complex (RISC) which further prevents selective mRNA expression. Researchers use this technique in order to selectively induce suppression of specific genes in order to examine how a cell functions without them (14). |
Figure 6: RNAi mechanism (15)
|
*This list is not exhaustive.
What is Gene Expression Analysis?
|
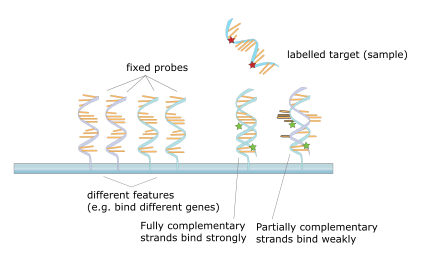
Figure 7: Sample microarray well (16)
|
While human cells have the potential to produce thousands of different proteins, they will actually only produce of a fraction at a given time. Gene expression analysis is a tool researchers use to study what kind of genes a cell is expressing and at what intensity they are being expressed. In order to accomplish this task, microarray technology has been developed to screen for thousands of gene at the same time. Microarrays are prepared by first inserting thousands of fixed genes into the small wells of a microarray (one gene per well). Flourescently labeled DNA probes are then added to the microarray. After several washes, flourescent probes that have been bound to the fixed genes will remain and their intensity and sequence can be determined. The more intense the emitted light, the stronger the DNA expression is within the well (17).
|
Known as Gene Expression Omnibus (GEO), this online database has been created to aid researchers by making microarray information very accessible. Scientists use this information to inform their own sequence analysis experiments and microarrays. The database is quite vast and contains sequence analysis for almost every studied gene (18).
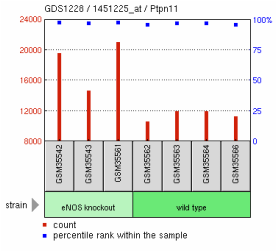
How do I interpret a GEO table?
1. Red bars show the measured number or value of an experiment.
2. Blue dots represent the percentile rank within the sample. These dots show a pattern that is important for understanding the trend the scientists found. Overall, this chart shows us eNOS knockout mice tend to have a higher rate of heart failure
3. Grey bars are the samples used in the study. This experiment had seven mouse samples.
4. Green bars show the conditions that each sample was subjected to. In this study the green bars were split into two groups: eNOS knockout and wild type.
Overall, PTPN11 is very well represented in the GEO database in terms of the diversity and number of studies available. The majority of the studies are unrelated to NS, but a few are related to symptoms presented by NS such as heart defects and body growth (19).
1. Red bars show the measured number or value of an experiment.
2. Blue dots represent the percentile rank within the sample. These dots show a pattern that is important for understanding the trend the scientists found. Overall, this chart shows us eNOS knockout mice tend to have a higher rate of heart failure
3. Grey bars are the samples used in the study. This experiment had seven mouse samples.
4. Green bars show the conditions that each sample was subjected to. In this study the green bars were split into two groups: eNOS knockout and wild type.
Overall, PTPN11 is very well represented in the GEO database in terms of the diversity and number of studies available. The majority of the studies are unrelated to NS, but a few are related to symptoms presented by NS such as heart defects and body growth (19).
References:
1) "PTPN11 Gene." Genetics Home Reference. N.p., n.d. Web. 27 Mar. 2015.
2) "Genes and Mapped Phenotypes." National Center for Biotechnology Information. U.S. National Library of Medicine, n.d. Web. 27 Mar. 2015.
3) "Gene Ontology Consortium." Gene Ontology Consortium. N.p., n.d. Web. 27 Mar. 2015.
4) "Kevin Ahern's Biochemistry (BB 450/550) at Oregon State University."Kevin Ahern's Biochemistry (BB 450/550) at Oregon State University. N.p., n.d. Web. 27 Mar. 2015.
5) Scott, Kyle. "Mechanisms of Hormonal Action." UTA (n.d.): n. pag. Web. 27 Mar. 2015.
6) Secko, David. "SCQ." SCQ. The Science Creative Quarterly, Aug. 2003. Web. 27 Mar. 2015.
7) "Human Aging Model Systems." : Cells, Yeast, Worms, Flies, and Mice. N.p., n.d. Web. 27 Mar. 2015.
8) "What Are 'model Organisms'?" What Are Model Organisms? N.p., n.d. Web. 27 Mar. 2015.
9) "ZFIN." ZFIN. N.p., n.d. Web. 27 Mar. 2015.
10) "Strain Detail Sheet." MMRRC:032104-JAX Detail Page (B6;129S4-Ptpn11tm1Bgn/Mmjax). N.p., n.d. Web. 27 Mar. 2015.
11) "Gene Dmel/csw." Flybase. N.p., 24 Feb. 2015. Web. 27 Mar. 2015.
12) "CRISPR/Cas9 Genome Editing." CRISPR Cas9 Genome Editing. N.p., n.d. Web. 27 Mar. 2015.
13) Sander, Jeffrey D., and J. K. Young. "CRISPR-Cas Systems for Editing, Regulating and Targeting Genomes." Nature. N.p., 2 Mar. 2014. Web. 27 Mar. 2015.
14) "RNAi." RNAi. N.p., n.d. Web. 27 Mar. 2015.
15) "RNAi Interference." Alnylam Pharmaceuticals. Alnylam Pharmaceuticals, n.d. Web. 27 Mar. 2015.
16) Huss, Mikael. RNA‐seq Analysis (2013): n. pag. 13 Feb. 2013. Web. 27 Mar. 2015.
17) "Gene Expression Analysis." Gene Expression Analysis. N.p., n.d. Web. 27 Mar. 2015.
18) "Getting Started." NCBI. U.S. National Library of Medicine, n.d. Web. 27 Mar. 2015.
19) "Ptpn11 - Heart Failure and Nitric Oxide." National Center for Biotechnology Information. U.S. National Library of Medicine, n.d. Web. 27 Mar. 2015.
Site Created by Daniel Turner - [email protected]
Updated on 5/16/15
www.genetics564.weebly.com
1) "PTPN11 Gene." Genetics Home Reference. N.p., n.d. Web. 27 Mar. 2015.
2) "Genes and Mapped Phenotypes." National Center for Biotechnology Information. U.S. National Library of Medicine, n.d. Web. 27 Mar. 2015.
3) "Gene Ontology Consortium." Gene Ontology Consortium. N.p., n.d. Web. 27 Mar. 2015.
4) "Kevin Ahern's Biochemistry (BB 450/550) at Oregon State University."Kevin Ahern's Biochemistry (BB 450/550) at Oregon State University. N.p., n.d. Web. 27 Mar. 2015.
5) Scott, Kyle. "Mechanisms of Hormonal Action." UTA (n.d.): n. pag. Web. 27 Mar. 2015.
6) Secko, David. "SCQ." SCQ. The Science Creative Quarterly, Aug. 2003. Web. 27 Mar. 2015.
7) "Human Aging Model Systems." : Cells, Yeast, Worms, Flies, and Mice. N.p., n.d. Web. 27 Mar. 2015.
8) "What Are 'model Organisms'?" What Are Model Organisms? N.p., n.d. Web. 27 Mar. 2015.
9) "ZFIN." ZFIN. N.p., n.d. Web. 27 Mar. 2015.
10) "Strain Detail Sheet." MMRRC:032104-JAX Detail Page (B6;129S4-Ptpn11tm1Bgn/Mmjax). N.p., n.d. Web. 27 Mar. 2015.
11) "Gene Dmel/csw." Flybase. N.p., 24 Feb. 2015. Web. 27 Mar. 2015.
12) "CRISPR/Cas9 Genome Editing." CRISPR Cas9 Genome Editing. N.p., n.d. Web. 27 Mar. 2015.
13) Sander, Jeffrey D., and J. K. Young. "CRISPR-Cas Systems for Editing, Regulating and Targeting Genomes." Nature. N.p., 2 Mar. 2014. Web. 27 Mar. 2015.
14) "RNAi." RNAi. N.p., n.d. Web. 27 Mar. 2015.
15) "RNAi Interference." Alnylam Pharmaceuticals. Alnylam Pharmaceuticals, n.d. Web. 27 Mar. 2015.
16) Huss, Mikael. RNA‐seq Analysis (2013): n. pag. 13 Feb. 2013. Web. 27 Mar. 2015.
17) "Gene Expression Analysis." Gene Expression Analysis. N.p., n.d. Web. 27 Mar. 2015.
18) "Getting Started." NCBI. U.S. National Library of Medicine, n.d. Web. 27 Mar. 2015.
19) "Ptpn11 - Heart Failure and Nitric Oxide." National Center for Biotechnology Information. U.S. National Library of Medicine, n.d. Web. 27 Mar. 2015.
Site Created by Daniel Turner - [email protected]
Updated on 5/16/15
www.genetics564.weebly.com